In DNA gene expression microarrays, thousands of gene expression levels are measured simultaneously. Microarray data may provide insight into gene to gene interactions, gene function, and pathway identification. The identification of differentially expressed genes is to find genes whose expression changes in response to different biological conditions. To identify genes potentially important in cancer, scientists have compared the global gene expression profiles of cancer tissue and corresponding normal tissue. Such analyses usually generate hundreds of genes differentially expressed in cancer relative to normal tissue, making it difficult to distinguish the genes that play a critical role in the neoplastic phenotype from those that are spuriously differentially expressed.
The core principle behind microarrays is hybridization between two DNA strands, the property of complementary nucleic acid sequences to specifically pair with each other by forming hydrogen bonds between complementary nucleotide base pairs. Total strength of the signal, from a spot, depends upon the amount of target sample binding to the probes present on that spot. Microarrays use relative quantization in which the intensity of a feature is compared to the intensity of the same feature under a different condition, and the identity of the feature is known by its position.
Bioconductor:
The analysis of microarray data requires biologists to learn the basics of statistics and programming. Many software tools for microarray data analysis are available. Currently one of the most popular and freely available software tools is Bioconductor. R Bioconductor is used to preprocess microarray data, detect differentially expressed genes, and annotate the gene lists of interest.
Prostate cancer:
Prostate cancer is a form of cancer that develops in the prostate, a gland in the male reproductive system. Most prostate cancers are slow growing; however, there are cases of aggressive prostate cancers.
Methodology:-
The main objective of this project is to identify new leads and target for Prostate cancer through Microarray data analysis and Structure based virtual screening which is a step in drug discovery.
Project involves two parts:
I-Microarray Data Analysis
II- Virtual Screening
In general, the methodology followed for accomplishing this project are as follows:
1- Microarray Data Analysis
1.1- Data collection from Gene Expression Omnibus(GEO-NCBI)
1.2- Normalisation in R(Bioconductor Package)
1.3- Statistical tests(T-test) using Multi Expression Viewer(MeV)
2- Lead and target identification
2.1- Building a network in cytoscape
2.2- Finding a good lead by screening library of compounds (Lipinki’s screening) and
and various online, offline softwares.
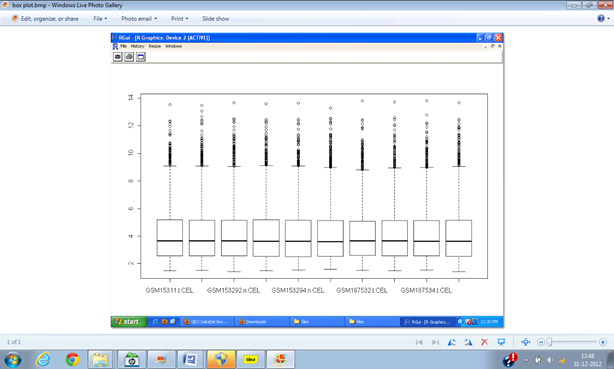
Graphs obtained from MeV:
Normalized graph-
Virtual screening (VS) is a computational technique used in drug discovery research. By using computers, it deals with the quick search of large libraries of chemical structures in order to identify those structures which are most likely to bind to a drug target, typically a proteinreceptor or enzyme. The aim of virtual screening is to identify molecules of novel chemical structure that bind to the macromolecular target of interest. There are two broad categories of screening techniques: ligand-based and structure-based. Given a set of structurally diverse ligands that binds to a receptor, a model of the receptor can be built by exploiting the collective information contained in such set of ligands.
And in Structure-based virtual screening which has been implemented in project, involves docking of candidate ligands into a protein target followed by applying a scoring function to estimate the likelihood that the ligand will bind to the protein with high affinity. The basic goal of the virtual screening is the reduction of the enormous virtual chemical space of small organic molecules, to synthesize and/or screen against a specific target protein, to a manageable number of the compound that inhibit a highest chance to lead to a drug candidate.
Glyceraldehyde 3-phosphate dehydrogenase (abbreviated as GAPDH ) is an enzyme of ~37kDa that catalyzes the sixth step of glycolysis and thus serves to break down glucose for energy and carbon molecules. In addition to this long established metabolic function, GAPDH has recently been implicated in several non-metabolic processes, includingtranscription activation, initiation of apoptosis, and shuttling, Metabolic function, as its name indicates, glyceraldehyde 3-phosphate dehydrogenase (GAPDH) catalyses the conversion ofglyceraldehyde 3-phosphate to D- glycerate 1,3-bisphosphate. This is the 6th step in the glycolytic breakdown of glucose, an important pathway of energy and carbon molecule supply which takes place in the cytosol of eukaryotic cells. The conversion occurs in two coupled steps. The first is favourable and allows the second unfavourable step to occur.
The first reaction is the oxidation of glyceraldehyde 3-phosphate at the carbon 1 position (the 4th carbon from glycolysis which is shown in the diagram), in which analdehyde is converted into a carboxylic acid (ΔG°'=-50 kJ/mol (-12kcal/mol)) and NAD+ is simultaneously reduced endergonically to NADH.
The energy released by this highly exergonic oxidation reaction drives the endergonic second reaction (ΔG°'=+50 kJ/mol (+12kcal/mol)), in which a molecule of inorganic phosphate is transferred to the GAP intermediate to form a product with high phosphoryl-transfer potential: 1,3-bisphosphoglycerate (1,3-BPG).
This is an example of phosphorylation coupled to oxidation, and the overall reaction is somewhat endergonic .Energy coupling here is made possible by GAPDH.
Generally, microarrays have expression levels for several thousand genes, and it is required to filter out those that are not significant. It is an important step before any further processing, such as clustering, to identify genes that are differentially expressed. The work focused on finding the best gene target, which can give better prediction accuracy for Prostate cancer. The best target as selected based on its role of cancer development was GAPDH i.e Glyceraldehyde 3-phosphate dehydrogenase. Finally, the work provides the best leads to the target by performing Virtual Screening of the natural anti cancer compounds.
Glyceraldehyde 3-phosphate dehydrogenase (abbreviated as GAPDH ) is the target as obtained after performing Mev and Cytoscape. Tripdiolide with molecular weight 376.400g/mol and Pomolic acid with 472.699g/mol are two leads which are suitable to bind to the target GAPDH in order to prevent the prostate cancer.
Web content composed with the free online HTML editor. Please purchase a membership to remove promotional messages like this.
References:
1. De Groot, A. S., et al. (2002). Immunology and Cell Biology, 80, 255â€"269.
2. Galperin, M. Y., & Koonin, E. V. (1999). Current Opinion in Biotechnology, 10, 571â€"578.
About Author / Additional Info:
M E in Bioinformatics