Authors: Pratik I. Bhoyar, Madhuri P. Suryawanshi and Amit Pippal
Introduction
Conventional plant breeding has been going on for hundreds of years, and is still commonly used today. Early farmers discovered that some crop plants could be artificially mated or cross-pollinated to increase yields. Desirable characteristics from different parent plants could also be combined in the offspring. When the science of plant breeding was further developed in the 20th century, plant breeders understood better how to select superior plants and breed them to create new and improved varieties of different crops. The conventional plant breeding is associated with some of its drawbacks like linkage drag, limited gene pool, and yield plateau. So to overcome the conventional plant breeding, biotechnological tools like genetic engineering and MAS (Marker assited selection) are widely used in addition to the conventional breeding tactices. The genetic engineering has the power to transfer gene from unrelated organism to the plants but it also has its limitation such biosafety issues and public acceptance. The need of new technology to develope better genotype is always confinding and the precise genome editing can sustain the need.
Genome Editing
Genome editing is the introduction of a predetermined sequence change to the chromosomal DNA of a cellular genome. The instructions for almost all functions of living systems are encoded in the genome. Consequently, the ability to easily and precisely add, remove, or exchange DNA sequences within a cellular genome would theoretically enable routine reprogramming of biological systems for numerous applications relevant to all areas of biotechnology, including medicine, energy, and the environment. The editing of genome sequences in diverse cell types and species has recently become possible through the advent of synthetic nucleases that can be engineered to target almost any site in a complex genome. The enhancement of gene targeting through nuclease-mediated DNA cleavage has been known for over fifteen years, but genome editing has not been widely applied to diverse areas of biotechnology until recently. This rapid growth is the result of the increased availability of public and commercial sources for engineering targeted nucleases, as well as significant progress in enhancing and monitoring genomic modifications. Despite the exponential growth of the use of this technology, current methods still do not fulfill the criteria of an ideal gene editing tool: (1) high frequency of desired sequence changes in the target cell population, (2) no off-target mutations, and (3) rapid and efficient assembly of nucleases that target any site in the genome at low cost.
Genome engineering is enabled by harnessing the cell’s DNA repair pathways. Most genome engineering techniques direct the repair of DNA double-strand-breaks (DSBs), which are introduced in the genome at or near the site where a DNA sequence modification is desired. Targeted DSBs are achieved using sequence-specific nucleases (SSNs) enzymes that recognize and cleave the target locus with high specificity. The repair of the break can be directed to create a variety of targeted DNA sequence modifications, ranging from DNA deletions to the insertion of large arrays of transgenes.
CRISPR/Cas system - Clustered regulatory intraspaced short palindromic repeats
A new tool based on a bacterial CRISPR-associated protein-9 nuclease (Cas9) from Streptococcus pyogenes has generated considerable excitement. This follows several attempts over the years to manipulate gene function, including homologous recombination and RNA interference (RNAi). RNAi, in particular, became a laboratory staple enabling inexpensive and high-throughput interrogation of gene function, but it is hampered by providing only temporary inhibition of gene function and unpredictable off-target effects. Other recent approaches to targeted genome modification â€" zinc-finger nucleases (ZFNs) and transcription-activator like effector nucleases (TALENs) enable researchers to generate permanent mutations by introducing doublestranded breaks to activate repair pathways. These approaches are costly and time-consuming to engineer, limiting their widespread use, particularly for large scale, highthroughput studies. The functions of CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) and CRISPR-associated (Cas) genes are essential in adaptive immunity in select bacteria and archaea, enabling the organisms to respond to and eliminate invading genetic material. These repeats were initially discovered in the 1980s in E. coli, but their function wasn’t confirmed until 2007 by Barrangou and colleagues, who demonstrated that S. thermophilus can acquire resistance against a bacteriophage by integrating a genome fragment of an infectious virus into its CRISPR locus. Three types of CRISPR mechanisms have been identified, of which type II the most is studied. In this case, invading DNA from viruses or plasmids is cut into small fragments and incorporated into a CRISPR locus amidst a series of short repeats (around 20 bps). The loci are transcribed, and transcripts are then processed to generate small RNAs (crRNA â€" CRISPR RNA), which are used to guide effector endonucleases that target invading DNA based on sequence complementarity. One Cas protein, Cas9 (also known as Csn1), has been shown, through knockdown and rescue experiments to be a key player in certain CRISPR mechanisms (specifically type II CRISPR systems). The type II CRISPR mechanism is unique compared to other CRISPR systems, as only one Cas protein (Cas9) is required for gene silencing. In type II systems, Cas9 participates in the processing of crRNAs, and is responsible for the destruction of the target DNA. Cas9’s function in both of these steps relies on the presence of two nuclease domains, a RuvC-like nuclease domain located at the amino terminus and a HNH-like nuclease domain that resides in the mid-region of the protein.
To achieve site-specific DNA recognition and cleavage, Cas9 must be complexed with both a crRNA and a separate trans-activating crRNA (tracrRNA or trRNA), that is partially complementary to the crRNA. The tracrRNA is required for crRNA maturation from a primary transcript encoding multiple pre-crRNAs. This occurs in the presence of RNase III and Cas9. During the destruction of target DNA, the HNH and RuvC-like nuclease domains cut both DNA strands, generating double-stranded breaks (DSBs) at sites defined by a 20-nucleotide target sequence within an associated crRNA transcript. The HNH domain cleaves the complementary strand, while the RuvC domain cleaves the noncomplementary strand. The double-stranded endonuclease activity of Cas9 also requires that a short conserved sequence, (2â€"5 nts) known as protospacerassociated motif (PAM), follows immediately 3´- of the crRNA complementary sequence. In fact, even fully complementary sequences are ignored by Cas9-RNA in the absence of a PAM sequence. system amenable to adaptation for genome editing. This potential was realized in 2012 by the Doudna and Charpentier labs. Based on the type II CRISPR system described previously, the authors developed a simplified two-component system by combining trRNA and crRNA into a single synthetic single guide RNA (sgRNA). sgRNAprogrammed Cas9 was shown to be as effective as Cas9 programmed with separate trRNA and crRNA in guiding targeted gene alterations.
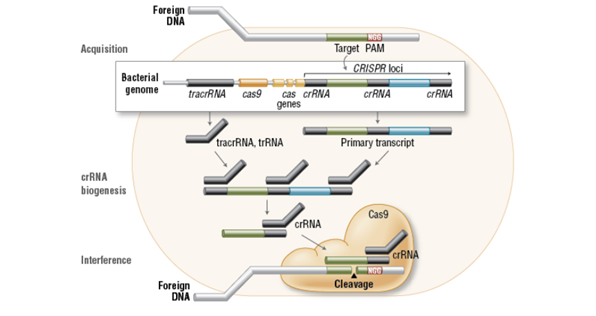
Fig: Mechanism of CRISPR
Regulation
How will plant varieties created through gene targeting be regulated? It is likely that each new variety will be considered on a case-by-case basis. The genetic variation that conferred resistance to late blight already existed in nature. Further, the resistant variety created through gene targeting is largely equivalent to a variety derived through traditional breedingâ€"an unregulated process. Since DNA-sequence variation closely linked to the resistance gene would also be introduced through breeding, gene targeting is actually more precise. The potato genome modified through gene targeting has only the desired DNA-sequence alteration, and this could be easily confirmed using approaches such as whole-genome sequencing. So the genotypes produced by genome engineering can surpass the regulatory process.
Conclusion
The genome editiong shows wide adaptability and capability for generating better genotype. The
improved genotype can sustain better agriculture which can be triggered with precise genome editing. The genome editing with CRISPER/Cas9 has its advantages over the othe nucleases like ZFNs and TALENs. It provide the possible way to edit the genome with precise modification for generation of the future genotypes.
References:
1. Gaj T., Gersbach C.A. and Barbas C.F. 2013. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends in Biotechnology. 31: 7.
2. Hartung F. and Schiemann J. 2014. Precise plant breeding using new genome editing techniques: opportunities, safety and regulation in the EU. The Plant Journal. 78: 742â€"752.
3. Nemudryi A.A., Valetdinova K.R., Medvedev S.P and Zakian S.M. 2014. TALEN and CRISPR/Cas Genome Editing Systems: Tools of Discovery. Acta Naturae. 6: 3 (22).
About Author / Additional Info:
I am currently pursuing Ph. D. in Molecular Biology and Biotechnology. I finished three years of Ph. D. and I am working on drought tollerent rice.