Authors: Jyoti Prakash Sahoo1* and Sandeep Kumar Singh2
Ph.D Research Scholar, 1Department of Agricultural Biotechnology, 2Department of Plant Breeding and Genetics, College of Agriculture, OUAT, Bhubaneswar - 751003
*Correspondence mail id : jyotiprakashsahoo2010@gmail.com
Introduction
Flooding, resulting in soil waterlogging and in many situations even complete submergence of plants, is an important abiotic stress in many regions worldwide. The number of floods has increased in recent decades and the severity of floods is expected to increase further in many regions of the world. Flooding reduces agricultural production, and floods shape many natural plant communities (e.g. floodplains, wetlands, salt marshes). Rainfed lowland and deep-water rice together account for approximately 33% of global rice farmlands (IRRI Social Statistics Database). Distribution of rice grown in upland, irrigated, rainfed lowland, and deep-water environments. Oftentimes, transient submergence is repeated or followed by a period of stagnant partial flooding. When partially or completely submerged, most rice varieties display a moderate capacity to elongate leaves and the portion of stems that are trapped underwater. This elongation growth leads to a spindly plant that easily lodges when floodwaters recede.
Early breeding of submergence tolerant rice
Landraces with unusual flooding and submergence tolerance were first reported in the early 1950s and systematically screened in the 1970s. The accessions FR13A and FR43B from Orissa, India and Kurkaruppan, Goda Heenati, and Thavalu from Sri Lanka were recognized for their resilience to complete submergence. FR13A stood out as extremely submergence tolerant; 100% of 10-day-old seedlings survived 7 days of complete submergence. However, FR13A lacks other agronomic attributes. It is photoperiod sensitive, tall, and provides low yields of poor-quality grain.
Mapping and molecular characterization of SUB1
Submergence of plants inhibits aerobic respiration and photosynthesis, and stimulates a variety of responses that can enhance survival, such as a switch from aerobic to anaerobic respiration. In contrast to deep-water rice cultivars that avoid submergence stress by growing above the water surface and thereby restoring gas exchange, submergence-tolerant rice can survive 10–14 days of complete submergence and renew growth when the water subsides, although the duration of survival is also influenced by environmental factors such as water turbidity, temperature and light levels. The Sub1 locus was mapped to an interval of 0.06 centimorgans on chromosome 9 using a mapping population (DX202) of 4,022 plants developed from the hybridization of a tolerant indica derivative of the FR13A cultivar (IR40931-26) and the intolerant japonica cultivar M-202 (Figure 1).
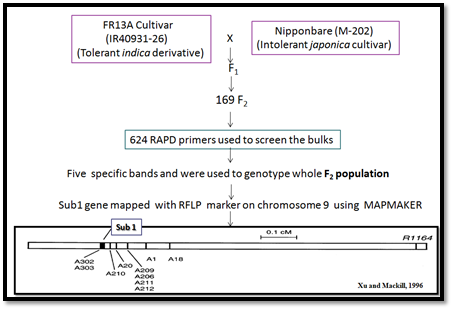
Figure 1. Mapping and molecular characterization of SUB1 locus
Genomic organization of Sub 1 QTL
The Sub1 region, borderedby themarkers CR25K and SSR1A, physically spans over 182 kilobases (kb). This interval encodes three genes containing ethylene response- factor (ERF) domains and designated Sub1A, Sub1B and Sub1C, ten non-ERF genes including four transcribed and six hypothetical protein-coding genes and 50% retrotransposon related sequences. The corresponding region of the japonica genome represented by the sequenced variety Nipponbare spans 142 kb and is considerably rearranged. SUB1A shares 87.7% and 77.2% sequence identity with SUB1B and SUB1C, respectively (Figure 2). Sub1A is absent from the Nipponbare genome. Recombination was suppressed in this region in the mapping population, as revealed by the 10.7-fold higher-than average recombination ratio (3,030 kb/cM in the Sub1 region versus 282 kb/cM for the entire genome). This could reflect the proximity of the Sub1 locus to the centromere and/or the presence of genomic rearrangements that have altered continuity in this region in the two rice subspecies.
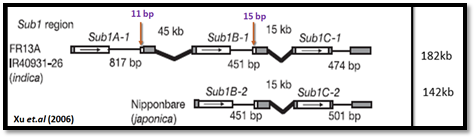
Figure 2. ERF gene structure and organization in tolerant indica (IR40931-26) and intolerant japonica (Nipponbare)
Haplotypes of the Sub1 locus
A haplotype is a set of DNA variation or polymorphisms that tend to inherited together. It also can refer to a set of single nucleotide polymorphisms (SNPs) found on the same chromosome. A survey of Sub1 locus haplotypes in 17 Indica and 4 Japonica varieties identified two Sub1A, nine Sub1B and seven Sub1C alleles on the basis of variation in AA sequence. Submergence tolerance is associated with the presence of Sub1 A-1, Sub1 B-1, 6, 3, Sub 1C-1 allele (Table 1).
Table 1. List of haplotypes of the Sub1 locus
| Line | Phenotype | Subspecies | Sub1A allele | Sub1B allele | Sub1C allele |
| FR13A | Tolerant | Indica | A1 | B-1 | C-1 |
| Goda Heenati | Tolerant | Indica | A1 | B-6 | C-1 |
| Kurkaruppan | Tolerant | Indica | A1 | B-3 | C-1 |
| IR64 | Intolerant | Indica | A2 | B-1 | C-3 |
| IR24, Swarna | Intolerant | Indica | Absent | B-8 | C-6 |
| IR50 | Intolerant | Indica | Absent | B-9 | C-7 |
| Nipponbare | Intolerant | Japonica | Absent | B-2 | C-2 |
| Liaogeng | Intolerant | Japonica | Absent | B-2 | C-2 |
| M-202 | Intolerant | Japonica | Absent | B-2 | C-2 |
Rescuing rice from flood
An Indian landrace Dhalputtia (FR13A), the source of the SUBMERGENCE1 (SUB1) locus that confers submergence tolerance by restricting underwater elongation growth, which is a quiescence strategy. Investigation of the functional role of the SUB1 genes determined that, as floodwaters rise, the gaseous phytohormone ethylene that is trapped within cells promotes SUB1A‑1 expression, which enhances the accumulation of two transcription factors that impede the response to gibberellin (GA): SLENDER RICE 1 (SLR1) and SLR1‑LIKE 1 (SLRL1). The outcome is a restriction of cell elongation and the less rapid consumption of energy reserves. Upon de‑submergence, genotypes with SUB1A‑1 incur less tissue damage from reactive oxygen species (ROS) and the dehydration that is associated with re-aeration. Even when leaf tissue loss is substantial, the maintenance of tiller meristems enables SUB1A‑1 genotypes to resume development when floodwaters subside. Submergence results in ethylene accumulation that activates transcription of the Sub1A gene. SUB1A represses Expansin A and sucrose synthase and, thus, represses growth. SUB1A also enhances mRNA accumulation and enzymatic activities of pyruvate decarboxylase (Pdc) and alcohol dehydroxygenase (Adh). Fermentation allows glycolysis to continue and, thus, provides the ATP needed for survival (Figure 3).
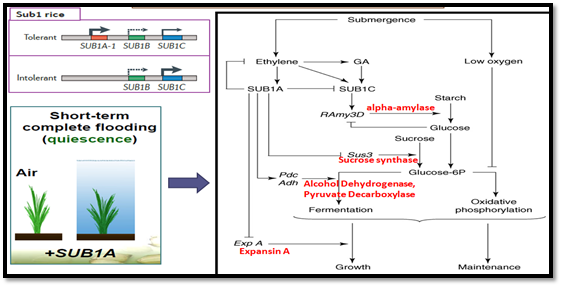
Figure 3. Mechanism of flooding survival in rice with Sub1
Introgression breeding of Sub1 mega varieties using Marker-assisted backcrossing (MABC)
MABC is a precise and an effective method to introgress a single locus controlling a trait of interest while retaining the essential characteristics of the RP. It is known that MABC is effective for genes or quantitative trait loci (QTLs) with large variations in phenotype. MABC is the process of using markers to select for target loci, minimize the length of the donor segment containing a target locus and/or accelerate the recovery of the RP genome during backcrossing. The main objective of MABC is to integrate a targeted gene from agronomical substandard sources (the donor parent) into an exclusive breeding line (the RP). MABC is superior to conventional backcrossing in precision and efficiency. The three selection steps are as follows (Figure 4).
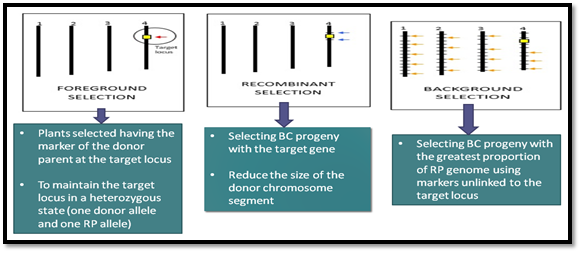
Figure 4. Marker-assisted backcrossing scheme for Introgression breeding of Sub1 mega varieties
Conclusion
The marker-assisted introgression of the SUB1 region has successfully improved submergence tolerance in a wide range of mega-varieties without any penalties on development, yield, and grain quality. These new lines endure submergence, as long as the flood occurs after the seedling stage but before flowering and the flood completely subsides within 10 to 20 days, depending on floodwater conditions. Although vegetative growth is restricted in some SUB1 varieties until the water level drops to 10–15 cm. Some recent studies showed that, this is not the case when SUB1 is transferred into taller varieties or those with better tolerance of partial stagnant flooding (20–50 cm). The yield advantage provided by SUB1 introgression lines is anticipated to greatly stabilize production in rainfed lowland environments that experience flash flooding.
References:
1. Septiningsih, E.M., Pamplona, A.M., Sanchez, D.L., Neeraja, C.N., Vergara, G.V., Heuer, S., Ismail, A.M. and Mackill, D.J. (2009). Development of submergence tolerant rice cultivars: the Sub1 locus and beyond. Annals of Botany 103: 151–160.
2. Xu, K., Xu, X., Fukao, T., Canlas, P., Maghirang-Rodriguez, R., Heuer, S. &Mackill, D. J. (2006). Sub1A is anethylene-response-factor-like gene that confers submergence tolerance to rice. Nature, 442: 705-708.
About Author / Additional Info: