Authors: Pratik I. Bhoyar, Madhuri P. Suryawanshi and Mahaveer
Introduction
The agriculture is affected by the wide range of factors. One of the main factor is genotype used. The agricultural research is directed towords production of the better genotype which can withstand present climatic condtion and perfom better. The conventonal breeding is slow in progress towords the aim so to complement the process biotechnological tools like MAS and transformation technology are used nowdays. The phenotyping is done in all experiememts as it help to justify the output of the experiments. So the precise phenotyping is the need of all experiments.
Conventional Phenotyping
The traditional phenotyping is acompanied with lots of bottlenecks to justify the experiments which cause the hinderence in the research programme and also time consuming and labour intensive. The whole phenotyping of the larger plot is quite difficult and if done which may be not precise. The traditional phenotyping is done by visual screeing which can not be useful for physiological and biochemical characteristic. Some of the biochemical charactarization includes destructive assay. The convetional phenotyping is error prone so the precise justification is difficult for the experiement. This is the reson of evolving the science of phenotyping.
Phenomics
The phenomics is the study of the phenome aiming to characterize phenotypes in a rigorous and formal way to link with the associated genes and gene variants (alleles). Plant phenomics deals with study of plant growth, performance and composition. Forward phenomics uses phenotyping tools to ‘sieve’ collections of germplasm for valuable traits. The sieve or screen could be high-throughput and fully automated and low resolution, followed by higher-resolution, lower-throughput measurements. Screens might include abiotic or biotic stress challenges and must be reproducible and of physiological relevance. Reverse phenomics is the detailed dissection of traits shown to be of value to reveal mechanistic understanding and allow exploitation of this mechanism in new approaches. This can involve reduction of a physiological trait to biochemical or biophysical processes and ultimately a gene or genes.
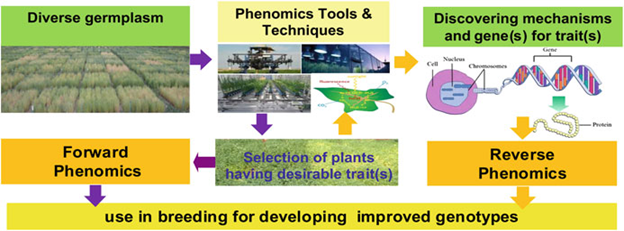
Figure: Diagramatic representation of the concept of phenomics
High-througput Phenotyping
The science of phenomics speeds up phenotyping by using automated high-tech sensors, imaging systems, moving vehicles and computing power. The main goal of phenomics is to bridge the gap between genomics, plant function and agricultural traits. The phenomics can help to overcome the bottleneck of the traditional phenotyping. The phenomics can be supported by the use of bioinformatics as the data generated will be huge. The associaiton of trait with genes is very precise with use of all such technologies under the diverse environmental condition.
Digital imaging can be used for obtaining measures of biomass and growth, and for probing some aspects of physiological function. Specific traits of particular interest to breeders, such as tillering, early vigor, coleoptile length and biomass at anthesis, can also be obtained from analyzed images. These traits are currently scored manually, and this can be improved in both speed and accuracy using phenomics tools. Proxies for some traits can be obtained by relatively simple properties of shape and texture, but the aim of image analyses is to develop 3-D models through time of the growing plants to enable extraction of more detailed, specific information, such as the life history of each leaf or even each tiller. This will allow the automated quantification of properties of particular interest of breeders working to improve traits underlying characteristics such as stress tolerance. For example, such models would allow the endurance of specific leaves to be quantified, which, in a salt-stressed plant, would allow the determination of the tolerance of the leaf to accumulated sodium ions (Na+).
Table: Imaging techniques used with respect to the trait to be phenotyped
| Sr. No. | Imaging Techniques | Traits to Phenotype |
| 1 | Visible Light | Shoot biomass , yield-related traits, leaf morphology, panicle traits, root architecture |
| 2 | Thermal Infrared | Leaf area index, shoot or leaf temperature, surface temperature, insect infestation of grain, leaf and canopy water status, |
| 3 | Hyperspectral | Water content, leaf growth and health status, panicle health status, grain quality, pigment composition, etc. |
| 4 | Fluroscense Imaging | Photosynthetic performance, quantum yield, non-photochemical quenching, leaf disease severity assessments, leaf health status, etc. |
| 5 | 3D imaging | Shoot structure; leaf angle distributions; canopy structure; root architecture; Height |
| 6 | MRI | Water content, morphometric parameters, etc |
| 7 | PET | Solute content, Metabolites content, etc |
The advantage of direct measurement of the trait of interest by imaging technologies rather than proxies is that it enables to exploit all the genetic variation responsible for a trait rather than one component. For example, one component of early seedling vigor, an important trait for conserving soil moisture, has been shown to be related to embryo size and is currently scored phenotypically by the width of the fully expanded leaves at young vegetative stage. Whether there are valuable sources of variation in vigor unrelated to this parameter has not been extensively examined because growth rate itself has not been amenable to high-throughput screening until recently.
The imaging techniques are accopanied with the moving vehicle to cover whole field. The phenocopter or high clearance tractors or drones can be used to generate the images. The genrated images are then analysed with software to findout the observation which are needed. The software platformas are also available with respect to part of the plant is to analyse. The bioinformatics adds the speed and accuracy in the phenotyping by managing the huge data and precise analysis.
Conclusion
Plant phenomics can considered as simply plant physiology in ‘new clothes’, but it promises to bring physiology up to speed with genomics by introducing the incredible recent advances made in computing, robotics, machine vision and image analysis to the wider field of plant biology. A multidisciplinary team in plant phenomics crosses biology, physics and mathematics, not ‘just’ genetics, biochemistry, physiology and plant breeding. This trans-disciplinary approach promises significant new breakthroughs in plant science. Phenomics provides the opportunity to study previously unexplored areas of plant science, and it provides the opportunity to bring together genetics and physiology to reveal the molecular genetic basis of a wide range of previously intractable plant processes.
References:
1. Cobb J.N., DeClerck G., Greenberg A., Clark R. and McCouch S. 2013. Next-generation phenotyping: requirements and strategies for enhancing our understanding of genotypeâ€"phenotype relationships and its relevance to crop improvement. Theor Appl Genet, 126:867â€"887.
2. Rahman H., Ramanathan V., Jagadeeshselvam N., Ramasamy S., Rajendran S., Ramachandran M., Sudheer P., Chauhan S., Natesan S., and Muthurajan R. 2015. Phenomics: Technologies and Applications in Plant and Agriculture. D. Barh et al. (eds.), PlantOmics: The Omics of Plant Science.
3. Rahaman M., Chen D., Gillani Z., Klukas C. and Chen M. 2015. Advanced phenotyping and phenotype data analysis for the study of plant growth and development. Frontier in plant science, 6:619.
About Author / Additional Info:
I am currently pursuing Ph. D. in Molecular Biology and Biotechnology. I finished three years of Ph. D. and I am working on drought tollerent rice.