Authors: Nimisha Tehri, Naresh Kumar, Raghu H.V., Pradip Kumar Sharma, Rajesh Gopaul and Geetika Thakur
1.0 - Introduction
The presence of various microbial an non-microbial contaminants as revealed from surveillance data available on food products has attributed to unhygienic environmental conditions during production, processing, transport, storage and indicates serious concerns. In recent years, wide spread outbreaks in various food products have reported to be linked with potential contaminants like melamine in China, pesticides in India, Listeria monocytogenes and Salmonella typhimurium in U.S.A. Keeping in view the findings of reports on massive outbreaks, limits of potential contaminants comprising of both microbial and non-microbial contaminants for different types of food matrices have been laid down by various regulatory bodies namely Codex Alimentarius, European Union, Food Safety and Standards Authority of India etc. This indicates the need for effective monitoring tools for targeting potential contaminants in various types of food matrices in order to comply regulatory standards. Till date, several conventional methods are available for detection of these contaminants. These methods are sensitive, efficient and reliable but have inherent drawbacks namely, limited scope of application under field conditions, time-consuming, and require complex pretreatment steps during estimation. Therefore, there is much demand for development of fast screening methods. Biosensors represent an interesting approach to detect various contaminants in food. They are analytical devices composed of two types of components which include biological recognition element and a suitable transducer. Biosensors as reported in prior art have been developed using enzymes, antibodies, aptamers, whole cells etc. as biosensing elements. The present article explores the use of bacterial spores as biorecognition molecules for detection of various types of contaminants in order to ensure food safety.
2.0 - Bacterial spores and their germination
Spores are dormant structure that are produced by a process of sporulation in few species of both Gram positive and Gram negative bacteria such as Bacillus, Clostridium, Sporomusa etc. to prevail over unfavourable environmental conditions. Being a survival strategy, endospores have very hardy and robust structure comprising of many layers such as core, inner membrane, germ cell wall, cortex, outer membrane, coat and sometimes exosporium as depicted in Fig.1. These structural properties of spores endow them with unique capability of preserving genetic material and to resist varying range of temperature, pressure, radiations, toxic chemicals, and many other extreme environmental conditions. Even though spores are resting bodies and formed under conditions of starvation and stress but they persistently scrutinize their external environment for various types of nutrients (sugars, amino acids etc.) and driving forces (temperature, pressure etc.) that allow them to germinate and thus converting them into metabolically active vegetative cells.
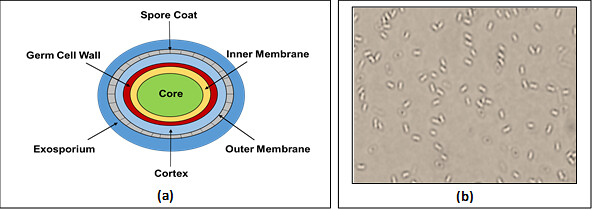
Fig. 1 (a) Schematic diagram of structure of bacterial spores (b) Dormant spores of a laboratory strain of Bacillus megaterium as observed under Phase contrast microscopy
3.0 - Spore as biorecognition molecule
The biphasic conversion of spores from dormant form to vegetative cell form and vice versa has been used as working principle for spores based sensors developed till date. These sensors employ the use of various biomarkers encompassing dipicolinic acid (DPA) and enzymes (e.g. proteases, lipases, esterases etc.) released during germination of spores. The presence of a particular analyte (contaminant) can signal the alteration of the process of germination which can be captured by using any of the aforementioned biomarker of germination.
4.0 - Spore based biosensors for different contaminants
Spores of different species of bacilli have been used as biosensing elements for development of biosensors in order to target various contaminants of microbial and non-microbial origin. The potential of developed spore based sensors have been evaluated by determining their suitability in terms of their application in analysis of contaminants in real food samples i.e. milk and milk products. Designing and analytical performances of developed sensors employing spores as biorecognition molecule for different types of contaminants are explained below.
4.1 - Detection of Antibiotics: Spores of Bacillus stearothermophilus have been used to develop biosensor for detection of antibiotic residues. The working principle of this sensor is based on the use of DPA as biomarker and a pH indicator. This sensor has successfully been applied for analysis of antibiotics in raw and pasteurized milk samples. The limit of detection (LOD) of this sensor is at or above maximum residues limit (MRL) established by regulatory bodies. The time of analysis is 3.0-3.15 hrs. [1]. Another biosensor has also been developed with spores of a different strain of B. stearothermophilus based on the principle of use of enzyme as biomarker for detection of antibiotic residues. The LOD with this sensor was also reported at or above MRL established by regulatory bodies but within lesser time of detection i.e. 2.0-2.15 hrs. as compared to DPA based spore biosensor [2].
4.2 - Detection of L. monocytogenes: A fluorimetric spore based sensor has been developed using spores of B. megaterium for detection of L. monocytogenes. The working of this sensor is based on the germinogenic substrate based concept. The LOD of this sensor with milk has been found to be 5.0 log cells using microtitre plate based assay and 3.0 log cells with biochip method using fluorescence reader and Electron Multiplying Charged Coupled Device (EMCCD) as a transducer respectively. The time of analysis is reported to be 4 hrs. (+ 12 hrs. enrichment) and 2 hrs. (+ 8 hrs. enrichment) with microtitre plate and biochip based assay respectively [3]. This sensor offer advantages in terms of less time requirement for analysis of samples as compared to conventional methods using a 7 days protocol.
4.3 - Detection of E. coli: A fluorimetric spore based sensor has been developed using spores of B. megaterium for detection of E.coli. The working of this sensor is based on the sugar uptake concept. The LOD of this sensor with milk has been found to be 5.0 log cells using microtitre plate based assay and 3.5 log cells with biochip method using fluorescence reader and EMCCD as a transducer respectively. The time of analysis is reported to be 2.30 hrs. (+6 hrs. enrichment) and 1.30 hrs. (+ 4 hrs. enrichment) with microtitre plate and biochip based assay respectively [4].
4.4 - Detection of Enterococci: A fluorimetric spore based sensor has been developed using spores of B. megaterium for detection of Enterococci. The working of this sensor is based on the germinogenic substrate based concept. The LOD of this sensor with milk has been found to be 5.6 log cells using microtitre plate based assay and 5.0 log cells with biochip method using fluorescence reader and EMCCD as a transducer respectively. The time of analysis is reported to be 2.30 hrs. (+ 10 hrs. enrichment) and 45 min. (+ 8 hrs. enrichment) with microtitre plate and biochip based assay respectively [5].
5.0 - Conclusions and future perspectives
Biosensors employing spores as biorecognition molecules for detection of different contaminants offer several advantages over existing conventional methods. For instance, spore based biosensing systems have been reported to be highly sensitive, reliable, user-friendly, cost-effective offering long shelf-life and on-site application for detection of contaminants. Thus using bacterial spores as biorecognition molecule holds great promises for targeting a broad range of contaminants such as pesticides, heavy metals, detergents, sanitizers, spoilage types and pathogenic microorganisms and several others contaminants important from public health point of view, in different types of food matrices in order to ensure food safety and quality assurance.
References
[1] Kumar, N., Sawant, S., Malik, R. K., and Patil, G.R. (2006). Development of analytical process for detection of antibiotic residues in milk using bacterial spores as biosensor. Indian Patent Reg No. 1479/DEL/2006.
[2] Kumar N., Khan A., Arora S., Patra F., Dhaiya M., Raghu H V., Balhara M., Sharma PK and Shaikh S (2014). Enzyme-Spore based assay (s) for detection of antibiotic residues in milk. Indian Patent no. 2213/DEL/2014.
[3] Balhara M., Kumar N., Thakur G., Raghu H. V., Singh V. K., Lawaniya R., Khan A. and Shabnam 2013. A Novel enzyme-substrate based bio-assay for real time detection of Listeria monocytogenes in milk. Indian Patent Reg No. 1357/DEL/2013.
[4] Kumar N., Lawaniya R., Avinash, Arora B., Vishweswaraiah H.R., Balhara M., Saurabh Kadyan and Vinai Kumar (2014). Marker enzymes and spore germination based assay for detection of E. coli in milk and milk products. Indian Patent no. 2214/DEL/2014.
[5] Kumar N., Kaur G., Thakur G., Raghu H. V., Singh N., Singh V. K. and Raghav N. (2012) Real time detection of Enterococci in dairy foods using spore germination based bioassay. Indian Patent Reg No. 119/ DEL/2012.
About Author / Additional Info:
I am pursuing Ph.D. in Dairy Microbiology from ICAR-National Dairy Research Institute, Karnal-132001 (Haryana), INDIA.